Programming and patterning of epithelial barriers
My laboratory, started in January 2022, studies how epithelial barriers are established during development to generate regionally specialized antimicrobial and immunomodulatory programs, and how these programs are maintained in adult tissues under homeostatic conditions. We focus on conserved principles by which epithelial identity is shaped by developmental cues and subsequently reinforced through interactions with commensal microbiota, systemic signals such as steroid hormones, and ongoing immune–epithelial crosstalk.
A central interest of the lab is understanding how epithelial differentiation across continuous tissue surfaces – such as the genitourinary tract – gives rise to spatially organized resident immune cell niches. We examine how gradients of epithelial gene expression – particularly chemokines and other immune-modulatory factors – pattern the localization, density, and functional specialization of tissue-resident immune cells at barrier surfaces exposed to the external environment. To address these questions, we combine mouse models and organotypic culture systems with spatial and single-cell transcriptomic approaches to resolve epithelial and immune heterogeneity at high resolution. We investigate how shared developmental origins can diverge into distinct barrier states along anatomical axes, providing a tractable framework to study how epithelial transcriptional programs coordinate local immune architecture.
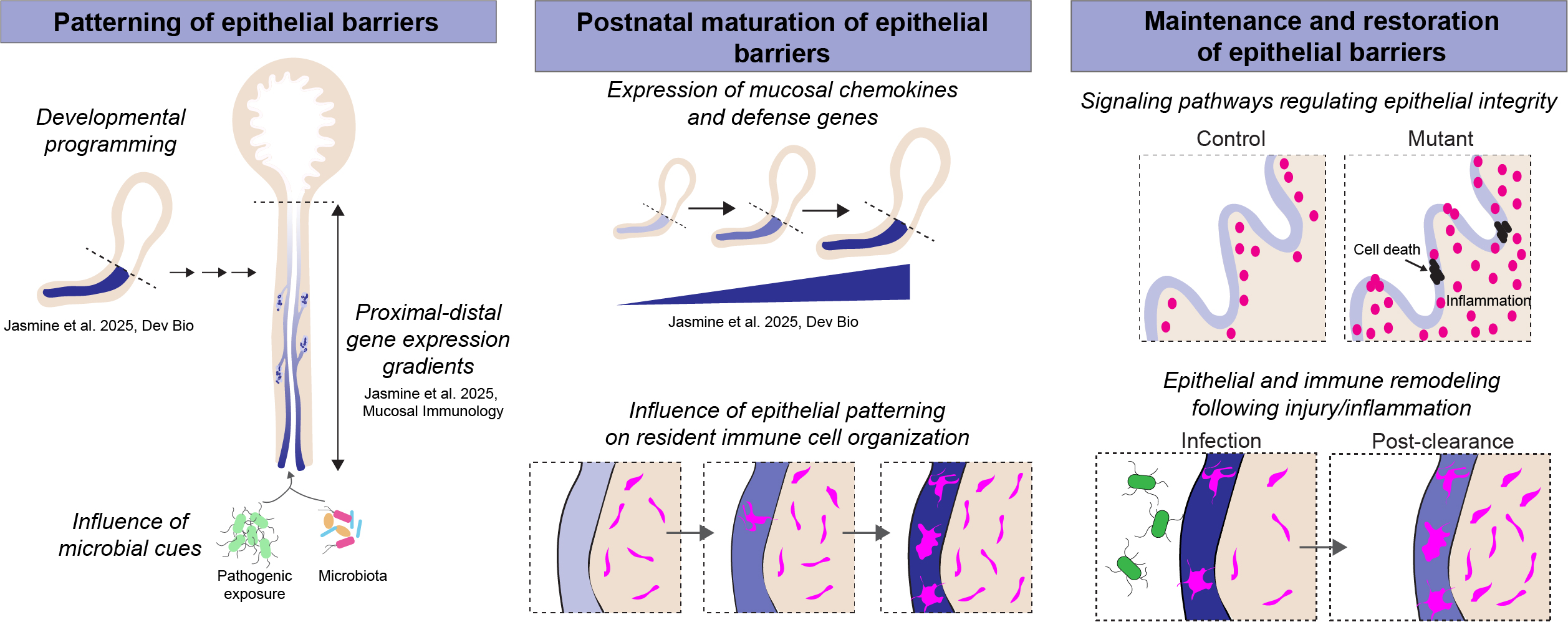
The bladder and urethra, with common developmental origins and contiguous epithelium, provide a useful model to study gene expression patterns along a proximal-distal axis. The urethral opening is an immunologically active site which has to contend with local microbiota and potential urinary pathogens. We identified that regional expression of barrier and chemokine genes in the urethral epithelium is established in utero and reinforced during postnatal development. (Jasmine et al. Developmental Biology, 2025). Our recent work (Jasmine et al. Mucosal Immunology, 2025) generated a spatial and single-cell atlas of the urethra, uncovering proximal-distal gradients in epithelial chemokine expression that correlate with spatial organization of epithelial-associated macrophages. These findings support a paradigm where temporal changes in epithelial gene expression organizes immune cell positioning to optimize barrier defense at sites of maximal environmental exposure.
Current efforts in the lab focus on defining how microbial-derived cues shape epithelial barrier maturation and resident immune organization in the genitourinary tract under homeostatic conditions and following repeated infectious or inflammatory challenges. We are investigating the role of signaling pathways like the NF-kB pathway in maintaining homeostasis in barrier tissues undergoing mechanical stress. Our new efforts are utilizing stem cell derived endometrial organoid models to understand the impact of metabolic rewiring of immune cells on epithelial homeostasis.
Research
Early epithelial patterning and establishment of immune defenses in the lower urinary tract
Epithelial barriers represent the first line of defense against an invasive pathogenic attack. Formation of epithelial barriers require patterning of contiguous epithelial tissue during development to generate specialized linings of organs. My first goal when establishing my program at inStem was to understand how gene expression in the urethra and bladder are patterned from the urogenital sinus and how immune defenses are established in the urethra during fetal development.
In this study, we provide a direct comparison of gene expression in the urethra and bladder during development, along with early transcription factors that are restricted to the bladder or urethra. We defined epithelial and mesenchymal gene signatures that are specific to the developing bladder or urethra. Previous studies have addressed bladder development in isolation from the urethra. By providing a direct comparison of gene expression in the epithelial and mesenchymal compartments of the lower urinary tract, we identified novel marker genes, transcription factors and signaling pathway genes that are spatially restricted to the bladder or urethra. Additionally, we identified a novel signature of chemokine gene expression in the urethral epithelium which correlated with the appearance of epithelial-associated macrophages. Lastly, our study revealed that the urethra shows sexually dimorphic gene expression in the epithelial and mesenchymal compartments while the bladder does not have the same degree of sexual dimorphism.
Publication: Jasmine, Divyeksha H. Baraiya, Kavya TT, Aparna Mandal, Shreya Chakraborty, Neha Sathish, Cynthia Marian Rebecca Francis, Diya Binoy Joseph*. Epithelial and mesenchymal compartments of the developing bladder and urethra display spatially distinct gene expression patterns. Dev Biol. 2025 Apr; 520:155-170. doi: 10.1016/j.ydbio.2025.01.005. PMID: 39798644.
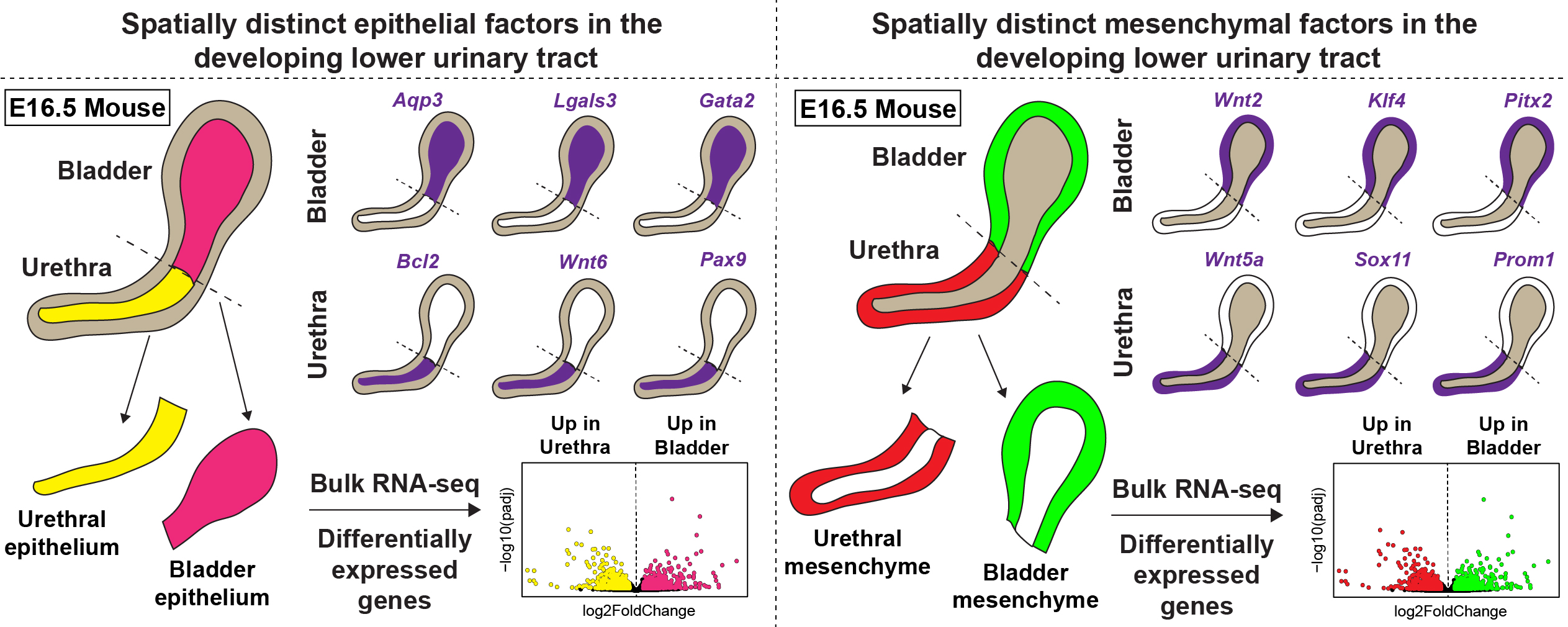
Identification of novel defense mechanisms in the female urethra
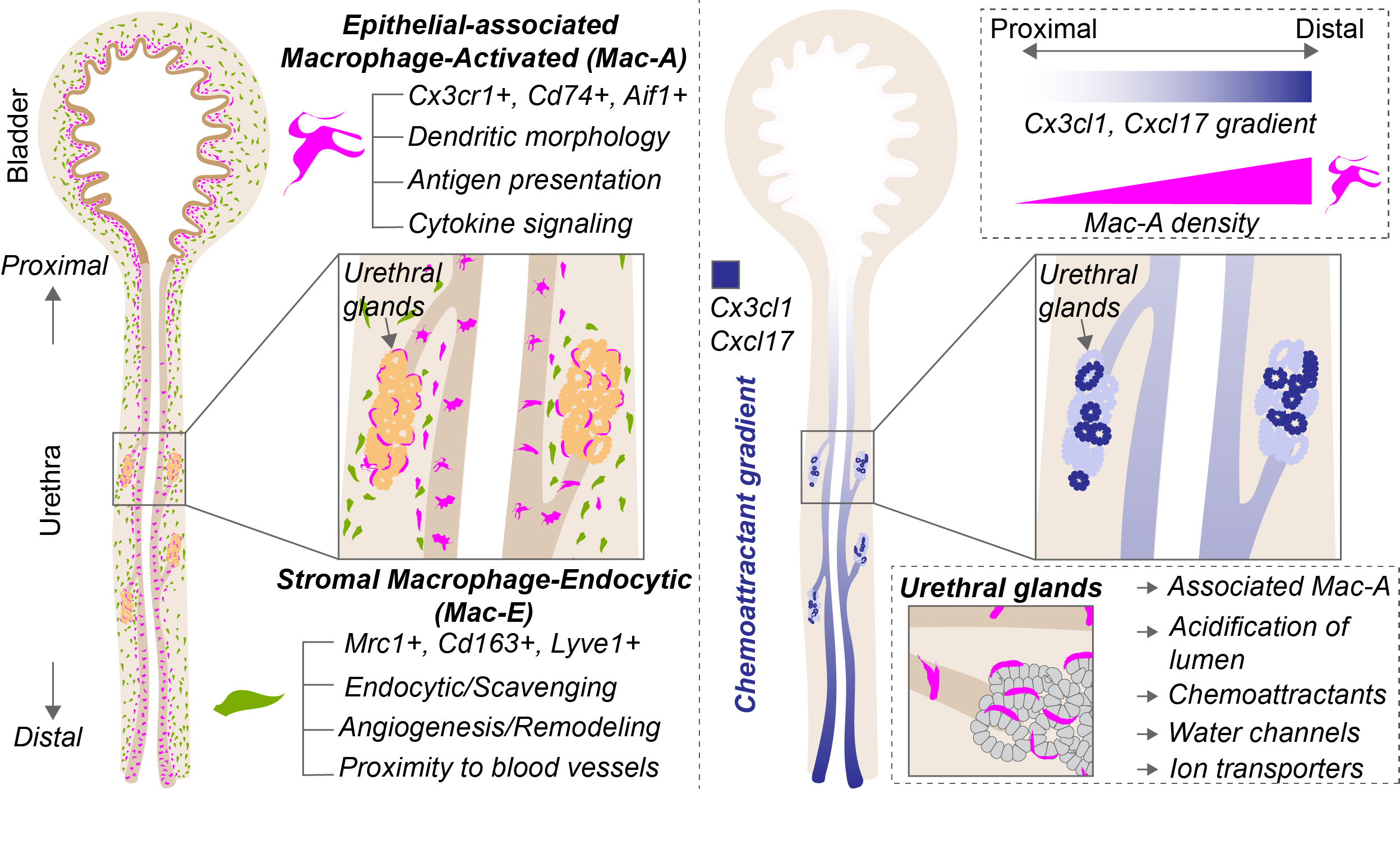
Urinary pathogens have to ascend the urethra to reach the bladder. The protective mechanisms in the urethra and how these are established are not known. We generated the first cellular atlas of the adult female mouse urethra revealing transcriptionally distinct basal, intermediate and luminal cells in the epithelial lining. Our work reveals the diversity of resident immune cell populations in the female urethra and identifies two spatially separated and transcriptionally distinct macrophage populations. One population of epithelial-associated macrophages displays a large cell perimeter and sends out dendrites in between epithelial cells. The other macrophage population resides in the stromal compartment and expresses gene signatures for scavenging, vascular development and tissue homeostasis. We also identify two secretory epithelial cell types in the adult female mouse urethra which express chemoattractant genes. Our work additionally uncovered novel urethral secretory glands which connected to the trunk of the urethra through short ducts. The urethral glands express antimicrobial genes and transporters for the acidification of lumen contents. Expression of chemoattractant genes in the urethral lining increases as we move closer to the outside of the body and correlates with the density of epithelial-associated macrophages which also increases towards the urethral opening. This study sheds light on an understudied mucosal barrier that is at the frontline of the response to urinary pathogens, identifying novel macrophage populations and epithelial gene signatures that can influence immunity.
Publication: Jasmine, Mrinal Samtiya, Richelle Rodrigues, Aparna Mandal, Kavya TT, Anubhuti Anushree, John T. Lafin, Chad M. Vezina, Douglas W. Strand, Diya Binoy Joseph*. Single cell map of the adult female mouse urethra reveals epithelial and stromal macrophages with distinct functional identities. Mucosal Immunology 2025 Sep 2: S1933-0219(25)00091-1. Epub ahead of print. PMID: 40907791.
Selected publications
PubMed Search terms: “Diya Binoy Joseph OR Diya Joseph”
NCBI Bibliography: https://www.ncbi.nlm.nih.gov/myncbi/1xMa5ZtM9NxAi/bibliography/public/
Google Scholar Profile: https://scholar.google.com/citations?user=YwwxE7IAAAAJ&hl=en
ORCiD Profile: https://orcid.org/0000-0002-0587-9558
Diya Binoy Joseph’s publications at inStem
- Jasmine, Mrinal Samtiya, Richelle Rodrigues, Aparna Mandal, Kavya TT, Anubhuti Anushree, John T. Lafin, Chad M. Vezina, Douglas W. Strand, Diya Binoy Joseph*. Single cell map of the adult female mouse urethra reveals epithelial and stromal macrophages with distinct functional identities. Mucosal Immunology 2025 Sep 2:S1933-0219(25)00091-1. Epub ahead of print. PMID: 40907791. *Corresponding author (SCIE, Impact factor 7.6)
- Jasmine, Divyeksha H. Baraiya, Kavya TT, Aparna Mandal, Shreya Chakraborty, Neha Sathish , Cynthia Marian Rebecca Francis, Diya Binoy Joseph*. Epithelial and mesenchymal compartments of the developing bladder and urethra display spatially distinct gene expression patterns. Dev Biol. 2025 Apr; 520:155-170. doi: 10.1016/j.ydbio.2025.01.005. Epub 2025 Jan 9. PMID: 39798644. *Corresponding author (SCIE, Impact factor 2.1)
Diya Binoy Joseph’s postdoctoral and PhD publications
- Diya B. Joseph, Gervaise H. Henry, Alicia Malewska, Jeffrey C. Reese, Ryan J. Mauck, Jeffrey C. Gahan, Ryan C. Hutchinson, James L. Mohler, Claus G. Roehrborn, Douglas W. Strand. ‘5-alpha reductase inhibitors induce a prostate luminal to club cell transition in human benign prostatic hyperplasia.’ Journal of Pathology (2022); 256(4), 427-441 PMID: 34928497 (SCIE, Impact factor 5.2)
- Diya B. Joseph, Gervaise H. Henry, Alicia Malewska, Jeffrey C. Reese, Ryan J. Mauck, Jeffrey C. Gahan, Ryan C. Hutchinson, Venkat S. Malladi, Claus G. Roehrborn, Chad M. Vezina, Douglas W. Strand. ‘Single cell analysis of mouse and human prostate reveals novel fibroblasts with specialized distribution and microenvironment interactions.’ Journal of Pathology (2021); 255(2), 141-154 PMID: 34173975 (SCIE, Impact factor 5.2)
- Diya B. Joseph, Gervaise H. Henry, Alicia Malewska, Nida S. Iqbal, Hannah M. Ruetten, Anne E. Turco, Lisa L. Abler, Simran K. Sand’hu, Mark T. Cadena, Venkat S. Malladi, Jeffrey C. Reese, Ryan J. Mauck, Jeffrey C. Gahan, Ryan C. Hutchinson, Claus G. Roehrborn, Linda A. Baker, Chad M. Vezina, Douglas W. Strand. ‘Urethral luminal epithelia are castration-insensitive cells of the proximal prostate.’ Prostate (2020); 80, 872-884 PMID: 32497356 (SCI, Impact factor: 2.5)
- Caden M. Ulschmid, Miranda R. Sun, Christopher R. Jabbarpour,….Ligia A. Papale, Diya B. Joseph, Chad M. Vezina, Reid S. Alisch, and Robert J. Lipinski. ‘Disruption of DNA methylation–mediated cranial neural crest proliferation and differentiation causes orofacial clefts in mice.’ Proc Natl Acad Sci U S A. (2024); 121(3):e2317668121 PMID: 38194455 (SCI, Impact factor: 9.1)
- Anne E. Turco, Steven R. Oakes, Kimberly P. Keil Stietz, Cheryl L. Dunham, Diya B. Joseph, Thrishna S. Chathurvedula, Nicholas M. Girardi, Andrew J. Schneider, Joseph Gawdzik, Celeste M. Sheftel, Peiqing Wang, Zunyi Wang, Dale E Bjorling, William A. Ricke, Weiping Tang, Laura L. Hernandez, Janet R. Keast, Adrian D. Bonev, Matthew D. Grimes, Douglas W. Strand, Nathan R. Tykocki, Robyn L. Tanguay, Richard E. Peterson, Chad M. Vezina. ‘A neuroanatomical mechanism linking perinatal TCDD exposure to lower urinary tract dysfunction in adulthood.’ Dis Model Mech (2021); 14(7):dmm049068 PMID: 34164643 (SCIE, Impact factor 3.3)
- Diya B. Joseph, Anne E. Turco, Chad M. Vezina, Douglas W. Strand. ‘Progenitors in prostate development and disease.’ Developmental Biology (2021); 473, 50-58 PMID: 33529704 (SCIE, Impact factor: 2.1)
- Su-Tang Lo, Daniel Parrott, M. Veronica Clavijo Jordan, Diya B. Joseph, Douglas Strand, U-Ging Lo, Ho Lin, Anza Darehshouri, A. Dean Sherry. ‘The roles of ZnT1 and ZnT4 in glucose-stimulated zinc secretion in prostate epithelial cells.’ Molecular Imaging and Biology (2020); 23, 230-240 PMID: 33140261 (SCIE, Impact factor: 2.5)
- Diya B. Joseph, Anoop S. Chandrashekar, Lisa L. Abler, Li-Fang Chu, James A. Thomson, Cathy Mendelsohn, and Chad M. Vezina. ‘In vivo replacement of damaged bladder urothelium by Wolffian duct epithelial cells.’ PNAS (2018); 115, 8394-8399 PMID: 30061411 (SCI, Impact factor: 9.1)
- Diya B. Joseph, Anoop S. Chandrashekar, Li-Fang Chu, James A. Thomson and Chad M. Vezina. ‘A folic acid-enriched diet attenuates prostate involution in response to androgen deprivation.’ Prostate (2018) ;79, 183-194 PMID: 30298636 (SCI, Impact factor: 2.5)
- Diya B. Joseph, Anoop S. Chandrashekar, Lisa L. Abler, Li-Fang Chu, James A. Thomson and Chad M. Vezina. ‘Epithelial DNA methyltransferase-1 regulates cell survival, growth and maturation in developing prostate buds.’ Developmental Biology (2019); 447, 157-169 PMID: 30659795 (SCIE, Impact factor: 2.1)
- Diya B. Joseph and Chad M. Vezina. ‘Male Reproductive Tract: Development Overview’ Encyclopedia of Reproduction 2nd edition, vol. 1, pp. 248–255, Elsevier 2018 (Book Chapter)
- Diya B. Joseph, Douglas W. Strand and Chad M. Vezina. ‘DNA methylation in development and disease: An overview for prostate researchers.’ Am J Clin Exp Urol (2018); 6(6),197-218 PMID: 30697577 (Impact factor: 1.4)
- Gervaise H. Henry, Alicia Malewska, Diya B. Joseph, Venkat S. Malladi, Jeon Lee, Jose Torrealba, Ryan J. Mauck, Jeffrey C. Gahan, Claus G. Roehrborn, Gary C. Hon, Jeffrey C. Reese, Ryan C. Hutchinson, Chad M. Vezina, Douglas W. Strand. ‘A Cellular Anatomy of the Normal Adult Human Prostate and Prostatic Urethra’ Cell Reports (2018); 25, 3530–3542 PMID: 30566875 (SCIE, Impact factor: 6.9)
- Sinoy S, Diya Binoy Joseph, Rajanikant GK. ‘Evolving Therapeutic Targets in Ischemic stroke: A Concise Review.’ Current Drug Targets (2013); 14(4), 497-506 PMID: 23278314 (SCIE, Impact factor: 2.5)
- Pradeep H, Diya Joseph, Shashikumar S, Rajanikant GK. ‘Oxidative Stress: Assassin behind the Ischemic Stroke’. Folia Neuropathol (2012); 50(3),219-30 PMID: 23023336 (SCIE, Impact factor: 1.6)
Lab members
Diya Binoy Joseph
I am a Scientist D and principal investigator at the Institute for Stem Cell Science and Regenerative Medicine (inStem) in Bengaluru. I have a degree in B. Tech. Biotechnology from the National Institute of Technology, Calicut, India. In 2018, I completed my PhD in Cellular and Molecular Biology from the University of Wisconsin-Madison, USA. My work involved using transgenic mouse models to look at the role of DNA methylation in lower urinary tract and prostate development. In 2019, I moved to UT Southwestern Medical Center in Dallas, USA for postdoctoral training where I was an American Urological Association Research fellow studying cellular heterogeneity and treatment response in Benign prostatic hyperplasia patients using single cell and spatial transcriptomics approaches.
I moved to inStem in January 2022 to start my independent lab. I am the recipient of a DBT-Wellcome Trust India Alliance Early career fellowship and a Start-up research grant from DST-SERB.
linkedin.com/in/diyabinoyjoseph/
Education and training
B. Tech Biotechnology, NIT-Calicut, 2013
PhD, Cellular and Molecular Biology, University of Wisconsin-Madison, 2018
Postdoctoral researcher, UT Southwestern Medical Center, Dallas, 2019-2021
Awards and distinctions
2023:
BDBio IISc, Big Data Algorithms for Biology, Best Post award (1 of 3 awarded)
2021:
American Urological Association (AUA) Annual Meeting Early-career investigator travel award
2020:
American Urological Association (AUA) Research Scholar Award 2020
Society for Basic Urologic Research (SBUR) virtual travel award
2019:
CMB Graduate Program Exceptional Thesis Award 2018, University of Wisconsin-Madison
Society for Basic Urologic Research (SBUR) travel award for podium presentation
2018:
CMB Graduate Program travel award, University of Wisconsin-Madison
2017:
Gregory F. Daniels award for conference travel, University of Wisconsin-Madison
Scholarship to attend Epigenomics course at Van Andel Research Institute, Grand Rapids
2016:
Society for Basic Urologic Research (SBUR) travel award for podium presentation
American Urological Association (AUA) Summer Research Conference travel award
2013:
Qualified GATE examination with All India Rank: 3 in Biotechnology (Score 984/1000)
‘Best Outgoing Student’ of NIT-C Class of 2013, Gold Medal
2012:
Khorana Scholarship, Dr. Arul Chinnaiyan group, University of Michigan, Ann Arbor
Qualified for CSIR-NET Junior Research Fellowship (JRF) with rank 29 in Life Sciences
Innovative project award NIT-C Alumni Association
Current lab members
Aparna Mandal, PhD Student (’22 batch)
Jasmine, PhD Student (’22 batch)
Mrinal Samtiya, Bridging Postdoctoral fellow
Amlan Banerjee, PhD Student (’24 batch)
Apoorva Roy, PhD Student (’25 batch)
Richelle Rodrigue, Project Associate II
Sharvari Vichare, Master’s dissertation student

Former lab members
Revanth Gowda, Master’s dissertation student
Tania Elizabeth Manoj, Summer intern
Surya K, Summer intern
Kavya TT, Project Associate I
Anubhuti Anushree, Summer intern
Arjun Suresh, Master’s dissertation student
Davood Irshad Meer, Master’s dissertation student
Divyeksha H Baraiya, Project Associate I
Shreya Chakraborty, Project Associate I
Cynthia Marian Elizabeth Francis, Project Associate II
Neha Satish, Summer Intern
Arshid Reshi, Master’s dissertation student
Hiring
We always welcome applications from interested candidates for internships and master’s thesis projects. Please write to diyabj@instem.res.in with a motivation statement and CV. For summer internships, please consider applying to the inStem Summer Research Fellowship Programme (SuRF). If interested in a PhD position in the laboratory, please apply through the inStem graduate student program or the BRIC-RCB i3c PhD Programme.



 Share
Share
 diyabj@instem.res.in
diyabj@instem.res.in 91-80-61948001 EXT, 8141
91-80-61948001 EXT, 8141
